RNA secondary structures play a crucial role in the function and evolution of RNA viruses, making them potential targets for therapeutic interventions. Identifying pivotal secondary structures is challenging, but artificial intelligence and machine learning techniques offer a promising approach to pinpoint these crucial domains. This approach could play a crucial role in managing and controlling future outbreaks of RNA viruses, such as SARS-CoV-2.
RNA viruses are among the most diverse and adaptable biological entities on Earth. They can rapidly evolve and infect new hosts, posing a significant threat to human health and well-being. One example of such a virus is the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which has caused the ongoing COVID-19 pandemic. SARS-CoV-2 is a single-stranded RNA (30 Kilobases long) virus that encodes both sequence and structure features that determine its function and interaction with host cells. However, these features are also subject to mutations that can affect the virus’s abilities such as its virulence and transmisibility. Some of these mutations have resulted in the emergence of new variants such as Delta, Omicron that have different characteristics and impacts on human health (Zabidi et al. 2023). Therefore, understanding the complexities of RNA viruses and their evolution is crucial for developing effective strategies to combat them. One critical aspect of understanding RNA viruses is studying the secondary structure of their RNA genome.
RNA Secondary Structure
Secondary structure refers to the folding of the RNA molecule into specific shapes and patterns, such as loops, stems, and hairpins, that are stabilised by base-pairing interactions. The secondary structure of RNA viruses influences their function and evolution, and play a vital role in many processes, such as replication, translation, packaging, and host recognition (Clyde and Harris 2006; Li et al. 2018). Therefore, utilising secondary structure information is crucial for a more accurate analysis of viral evolution as well as develop advanced therapeutics.
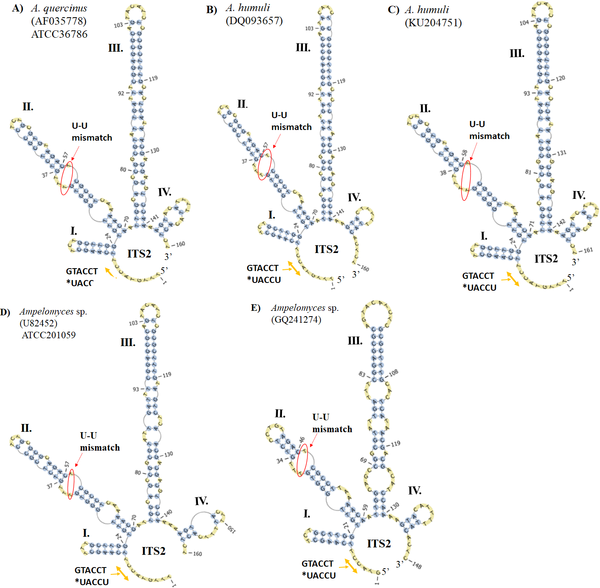
Incorporating Secondary Structure for Viral Evolution Analysis
Phylogenetic analysis, which compares the genetic sequences of different viruses to construct relationships, can reveal the origin, spread, and unique features of various viral variants. However, phylogenetic analysis based on DNA evolutionary models may not capture the full complexity of RNA viruses, due to their secondary structures that influence their function and adaptation. Therefore, RNA evolutionary models that incorporate secondary structure data can provide more accurate results (Patiño-Galindo et al. 2018).
However, one of the challenges of using RNA evolutionary models is the lack of secondary structure data for many RNA viruses. Secondary structure data can be obtained from experimental methods, such as selective 2’-hydroxyl acylation analysed by primer extension (SHAPE). However, these methods are either costly, time-consuming, or dependent on the quality and quantity of the input sequences. Therefore, there is a need for more efficient and reliable ways to generate and validate secondary structure data for RNA viruses.
One possible solution is to use machine learning methods that can predict secondary structure from sequence data such as the recent tools, SPOT-RNA and e2efold that utilise deep learning framework (convolutional neural networks) to increase structure prediction performance.
Targeting RNA Secondary Structures hotspots for interventions
Previous research has shown that RNA viruses adopt various conformations to establish specific base-pair interactions with the host's RNA (Guo and Steitz 2014). These interactions are pivotal for the virus's adaptability and virulence. Disrupting the secondary structure of a key genomic domain linked to virulence, using small interfering RNA (siRNA), for instance, could make the virus less adaptive to its host environment. Even though genomic mutations may persist, destabilising a crucial secondary structure could make the virus less viable, ultimately leading to its elimination through natural selection mechanisms (Prahl et al. 2023).
Yet, a significant challenge remains: how can we pinpoint which structures to target and their locations, for example, in the nearly 30 Kb genome of SARS-CoV-2? Although the answers remain elusive, artificial intelligence and machine learning techniques, such as deep neural networks, offer a promising approach to identifying the pivotal structural domains essential for viral function and virulence.
While there are ample databases and resources for modelling protein structure, function, and their interrelationships, RNA resources are considerably fewer. Leveraging machine learning can speed up the process of uncovering relationships between structure and function. However, building a machine learning model requires a significant amount of data to uncover previously unidentified patterns (Sarker 2021). Transfer learning can circumvent the dataset size limitation by relaxing the requirement for training and test data to be independent and identically distributed (Sun et al 2019).
By training a novel algorithm on existing data that document known RNA modifications—both in sequence and structure—that lead to functional disruptions, we can swiftly identify potential RNA domains crucial for virulence. These identified regions could then be deemed 'hot spots' for potential therapeutic intervention. Nonetheless, it is crucial to rely on experimentally validated secondary structures.
Conclusion
In summary, developing a new computational tool based on machine learning will assist us in identifying motifs with structural functions, such as replication, virulence, or other viral traits linked to pathogenicity. Subsequently, we can determine the likelihood of the 'hit' structure-function relationship being affected. Although viruses will continue to mutate and potentially evade the immunological response triggered by targeted vaccines, it is unlikely they will escape the structural constraints imposed by disrupting the 'hit' structure. This is because more than one simultaneous mutation may be required for that purpose. In this context, viruses will be selectively eliminated despite mutations. While we have vaccines that protect against severe viral illness, our new approach will enable us to reduce the replication of new circulating variants within the host. Ultimately, this approach represents a promising new frontier in the fight against RNA viruses and could play a crucial role in managing and controlling future outbreaks.
References
Subscribe to Transformational Bioinformatics
Stay up-to-date with our progress