An overwhelming amount of data is being generated in every thriving field today. Specifically, the amount of information per sample is growing exponentially every year especially in the field of Bioinformatics. It has been common practice for bioinformaticians to reduce the size of data first and then perform analysis.
However, this approach is not ideal as every bit of data can provide information. Things that seem irrelevant now might later become relevant as more data gets processed. Such data reduction strategies might prove to be particularly dangerous as they could lead to incomplete or incorrect biological insights.
Importance of choosing the right compute architecture
Advances in compute technology make this flawed approach outdated: With the correct database and data architecture, almost all current biological big data processing needs can be addressed. Just like one key is not a fit for multiple locks, so can one database solution not cater to all types of data needs.
It is very important to take into account factors such as data type, data growth, data retrieval modes, and data retrieval frequency when designing the compute architecture at the very start of a project.
Choosing the right data architecture is like building a solid foundation at the start of a construction: the stronger and more planned the architecture the greater the scope to expand the application later.
We are also at the physical limits of being able to increase the processing power on single CPUs. Hence the free increase in compute power we've experienced with every generation of processor is exhausted. Traditional high performance compute approaches require larger and larger clusters to cater for the demand increase. This in turn is not sustainable for individual organisations.
Cloud as a solution for biological applications
Cloud compute was the obvious answer for us. Creating a customized computing cluster, based on your specific (hardware) needs and paying only for what you used, was far-fetched 12-15 years back but it is a reality now. Public cloud platforms are offering a range of data architecture and computing solutions that can be combined to provide the ideal setup for the required analysis.
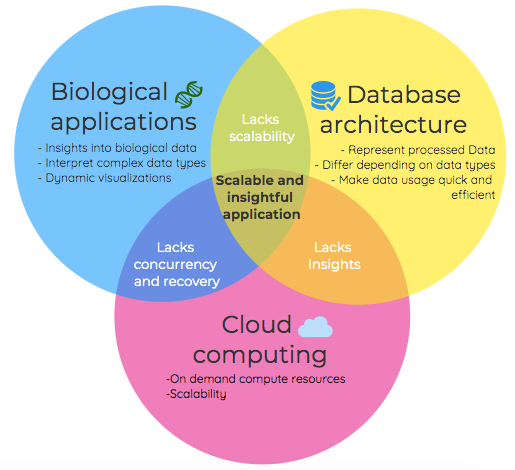
We, therefore, argue that future biological projects required domain knowledge, the elasticity of cloud computing and efficient data access to create scalable and insightful applications of the future.
VariantSpark: an example of scalable and insightful bioinformatics
VariantSpark is one such pioneering example of how accurate biological insights could be generated without biasing or compromising input data and volume. VariantSpark is built on top of Apache Spark – a modern distributed framework for big data processing, which gives VariantSpark the ability to scale horizontally.
Biological software solutions need to be flexible enough to adopt any cloud technology to cope with the increased sizes of biological datasets. It is also important that software is extensible to store the generated data to produce more insights and visualizations.
VariantSpark demonstrates this flexibility as it is currently available on top 3 major cloud services providers: Amazon Web Services (AWS), Microsoft Azure and Google Cloud Platform (GCP) and taps into established technologies for its visualisation (Cytoscape).
Where to from here?
VariantSpark could also be extended in the future to work with modern NoSql databases like Riak KV, which is one of the first to also utilize Apache Spark. Keeping both data access and compute on distributed compute architecture allows novel advances in terms of speed as well as other domains such as data security.
If you want to find out more about any of these topics get in touch.


